| Team Members: | Kallista Angeloff1, Carlos Barajas2, Alexander D. Middleton3, and Uchenna Osia4 |
| Graduate Assistant: | Jonathan S. Graf5 |
| Faculty Mentor: | Matthias K. Gobbert5 |
| Client: | Zana Coulibaly6 |
1Department of Atmospheric Sciences, University of Washington,
2Department of Mathematics and Computer Science, Olivet College,
3Department of Mathematics, Winthrop University,
4Department of Computer Science and Electrical Engineering, University of Maryland, Baltimore County
5Department of Mathematics and Statistics, University of Maryland, Baltimore County,
6Department of Pharmacology, University of California, Davis
About the Team
Our team – Kallista Angeloff, Carlos Barajas, Alexander D. Middleton, and Uchenna Osia – extended a partial differential equations model of calcium induced calcium release (CICR) in cardiomyocytes to complete the links between the electrical, chemical, and physical dynamics involved. Our model was written in C and fully parallelized using MPI on the HPCF cluster maya. We conducted our research in the summer of 2016 at the UMBC REU Site: Interdiciplinary Program in High Performance Computing, with guidance and assistance from our faculty mentor Matthias K. Gobbert and our graduate assistant Jonathan S. Graf.
Problem
Calcium dysregulation is a significant cause of fatal cardiac arrythmias, but it is an incompletely understood phenomenon and difficult to predict. Heartbeat rhythm is governed by periodic membrane depolarizations, causing the release of Ca2+ ions into the cytosol of individual cardiomyocytes; the reaction of this calcium with contractile proteins triggers the overall contraction of the heart. These calcium patterns can be modelled as a system of coupled partial differential equations linking the excitation, signaling, and contraction of individual cardiomyocytes.
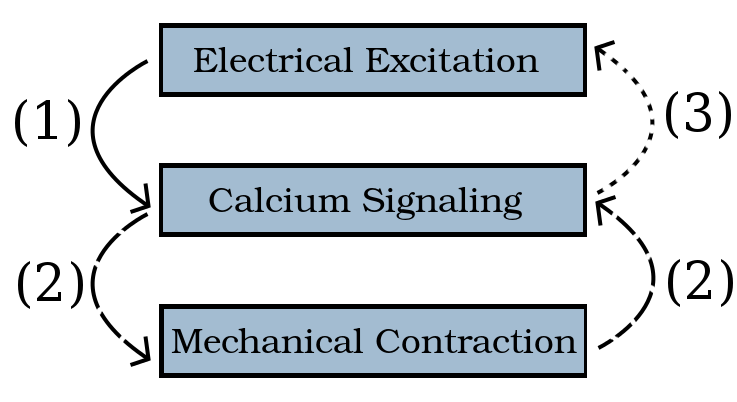

The starting point of this research, the work of 2015’s Team 5, was a model which established a one-way connection ① from the electrical to the chemical systems of the cell. Our work ②, ③, ④ extends this earlier implementation to complete the model and to couple all three systems fully. The initial simulations reported here compare one-way coupling ① to two-way coupling ①–② of the electrical and calcium systems.
Methodology
Building on code developed in C, we extended the model to link all the dynamics involved in a heart cell. Our first addition was a parameter to couple the chemical system to the electrical system via the current generated by calcium efflux; this current, dependent on cytosol calcium concentration, was built into the term governing membrane potential to make the voltage calcium-dependent. Our second addition was a cytosol buffer species for a contractile protein complex: we simulated cell contraction by modelling the changing proportion of these complexes in reaction with calcium and troponin, another contractile protein. After formulating these parameters and reaction terms, we conducted a parameter study to determine the most physiologically realistic outcomes of the model across the range of our feedback strength ω.
Results
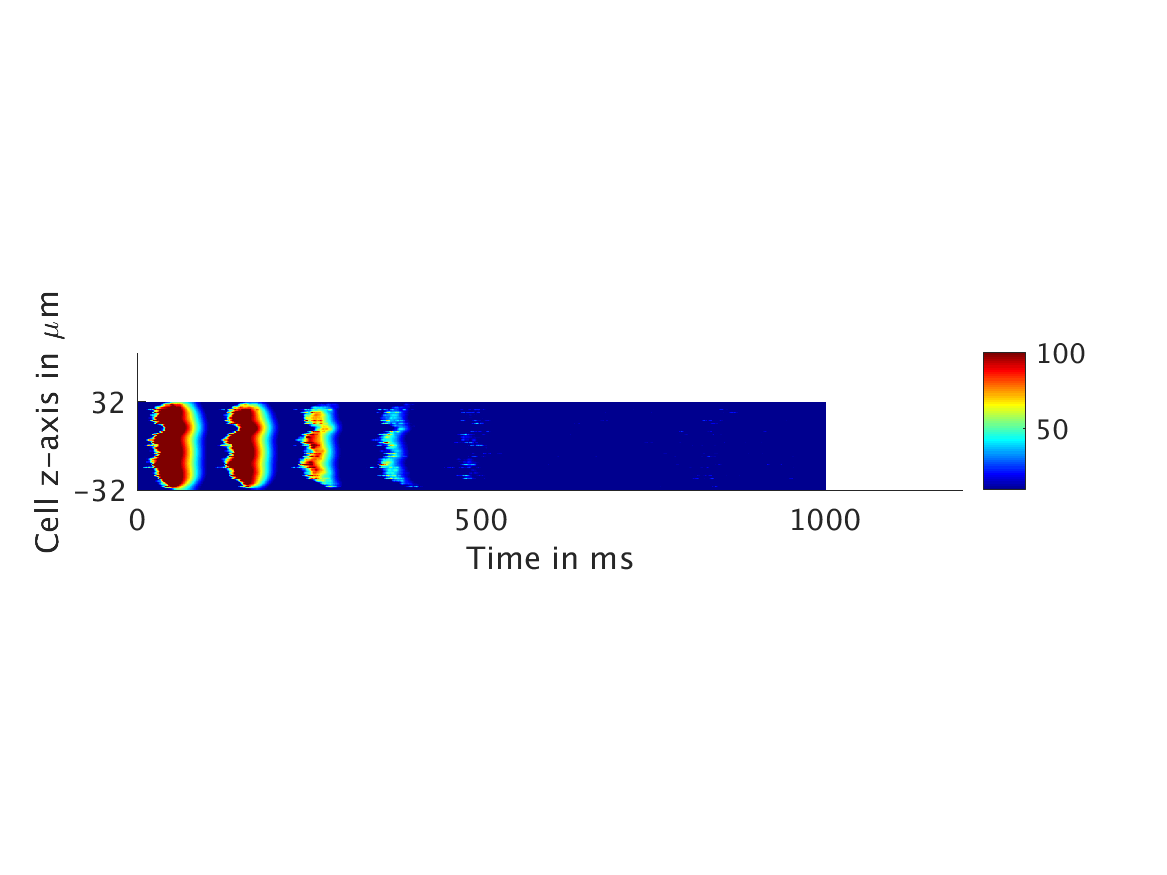
We present two different types of plots. In the left of each figure, a two dimensional line scan image of the calcium concentration in the cytosol with dark blue indicating low concentration of calcium and dark red indicating high concentration of calcium. On the right half of each figure are voltage plots versus time at the center of the cell.
Shown here is the one-way coupling ①, with
ω = 0:
[***CAPTIONS: Line scans /// Voltage vs. t***]

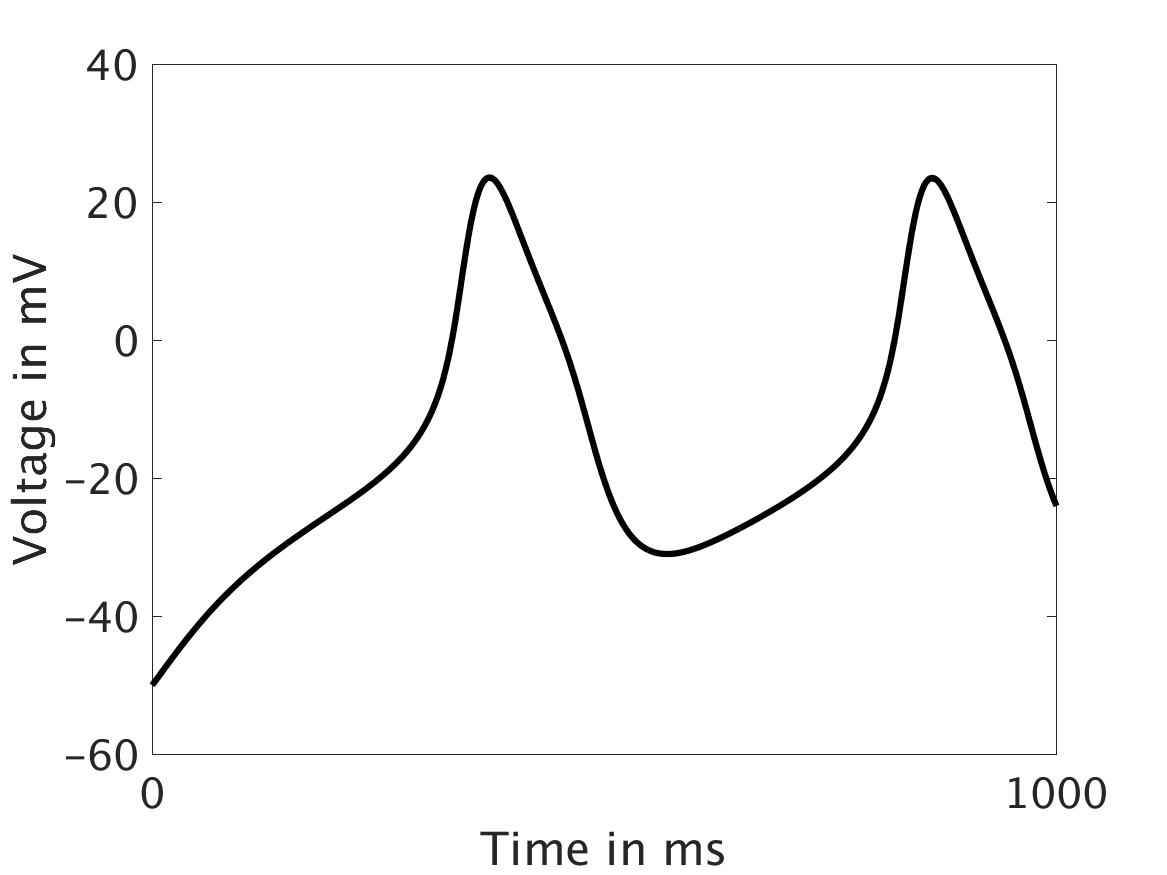
The voltage is unaffected by the fluctuations in calcium concentration. This is not physiologically realistic.
We then tested low values of ω and observed that the feedback strength was too low to have meaningful influence in the voltage:
ω = 0.01:

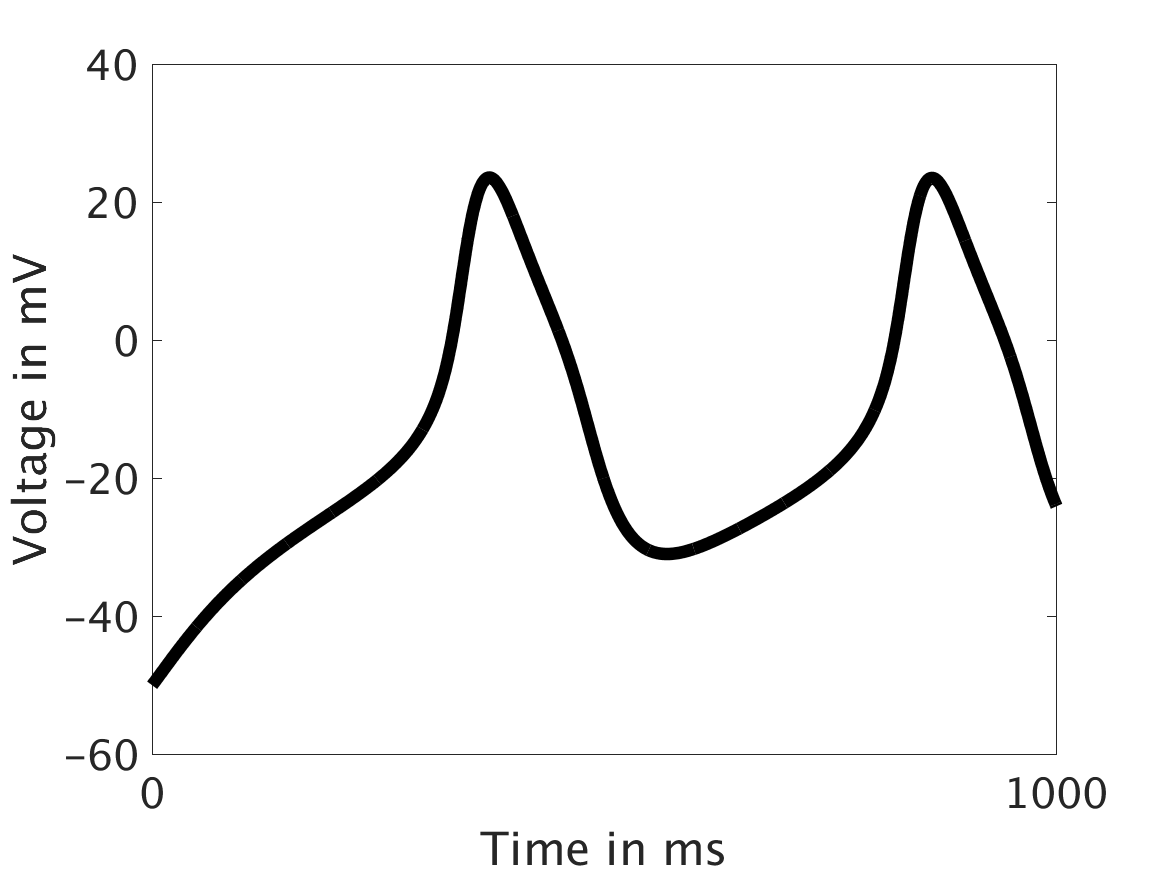
ω = 0.1:
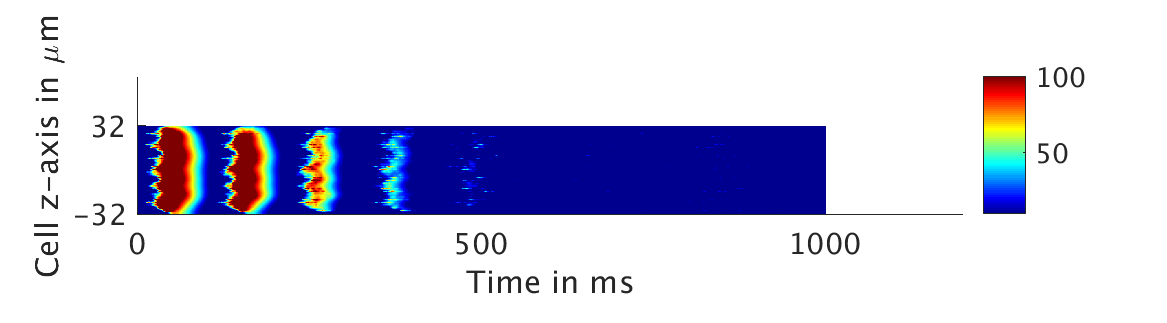

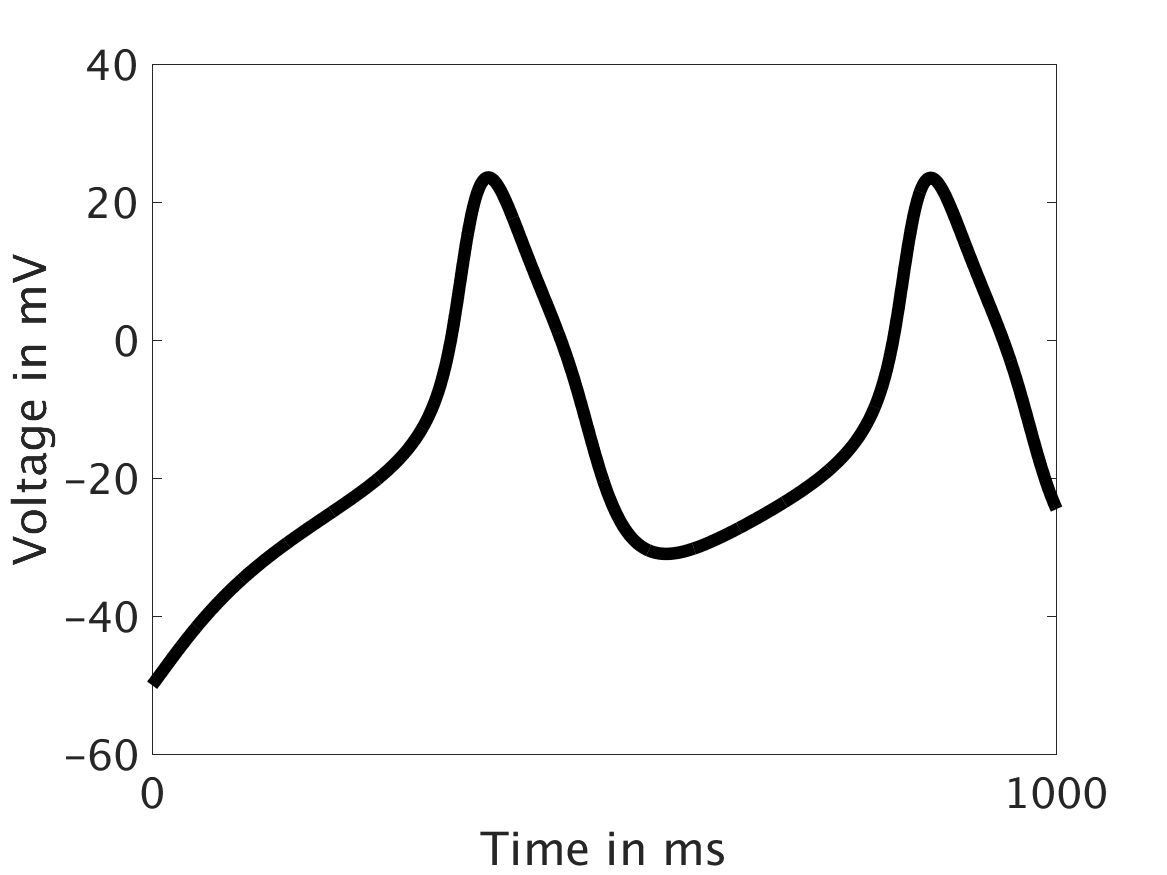
ω = 1:
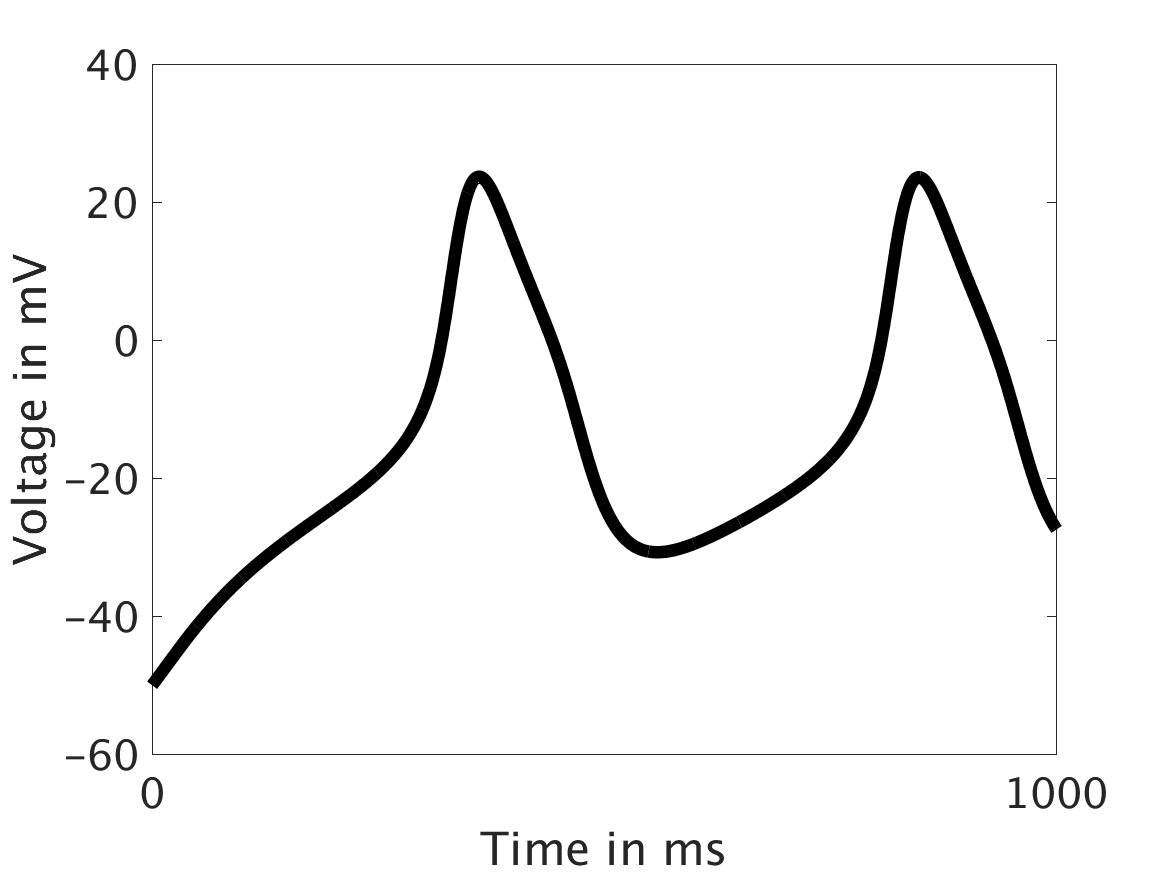
However, as we increased ω and correspondingly increased the effect of calcium concentration on the voltage, we observed that the formerly periodic behavior of the voltage was more realistically destabilized by the sharp fluctuations in current caused by the calcium oscillations.
ω = 30:
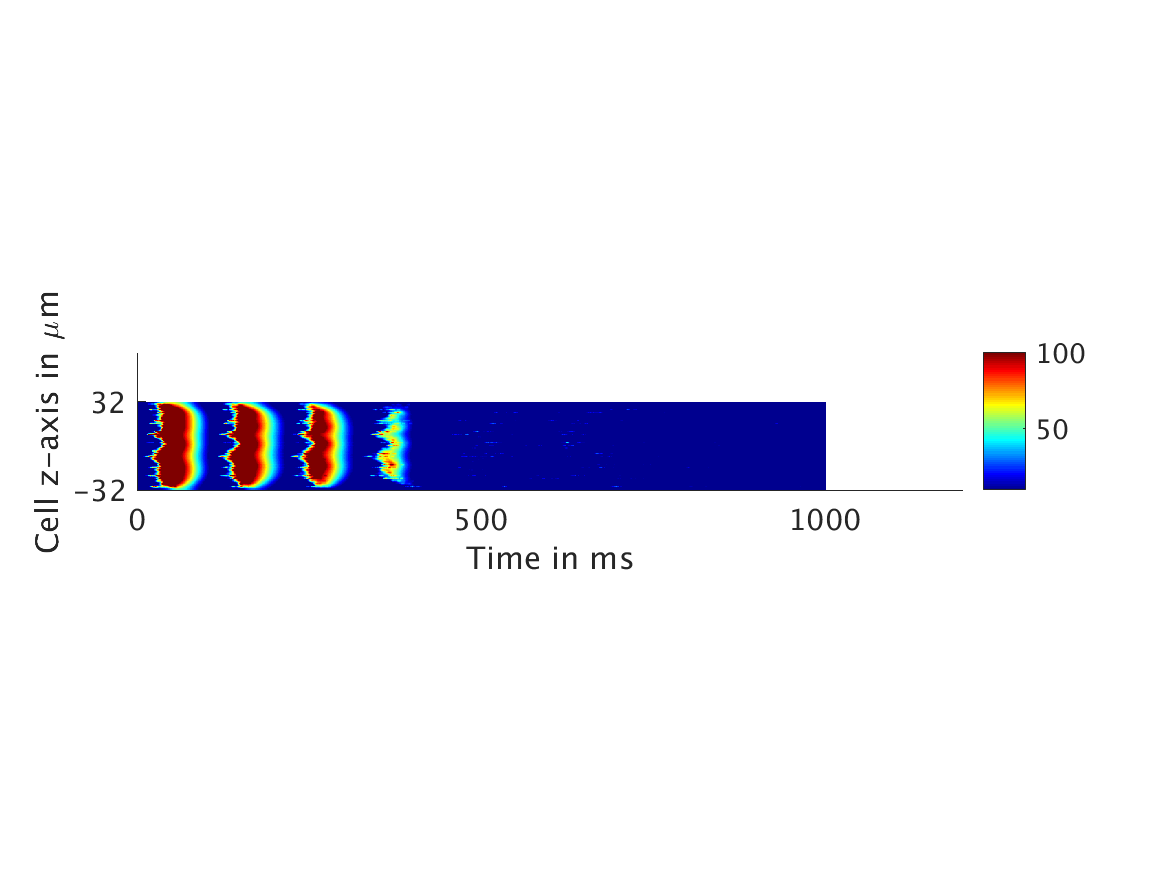
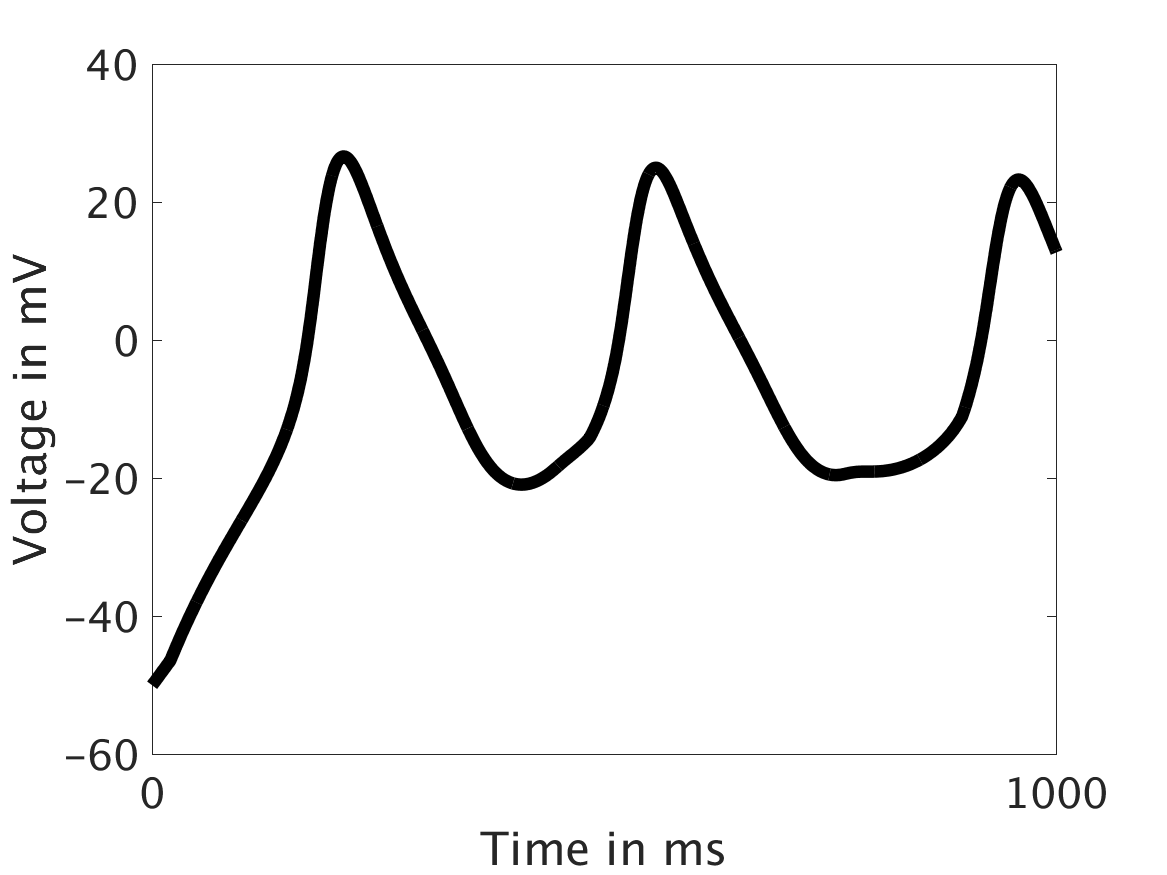
ω = 50:
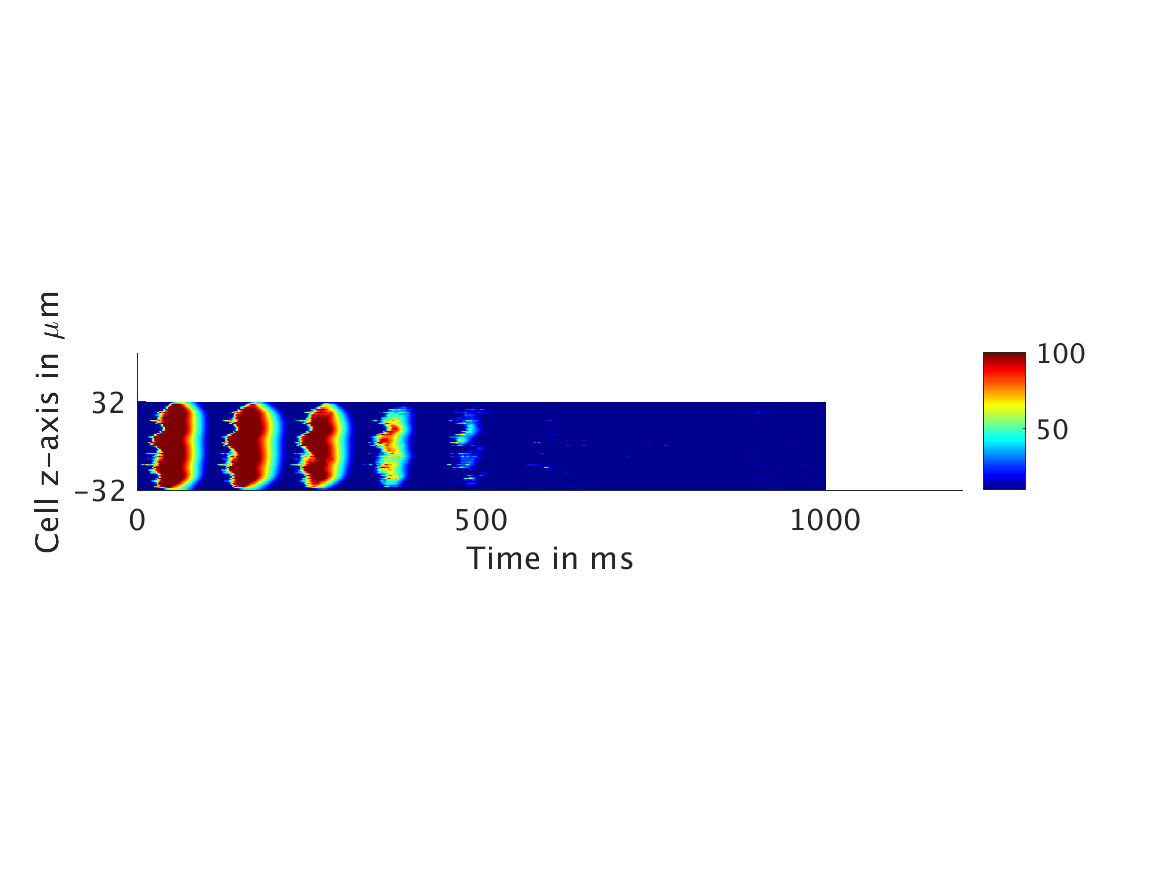
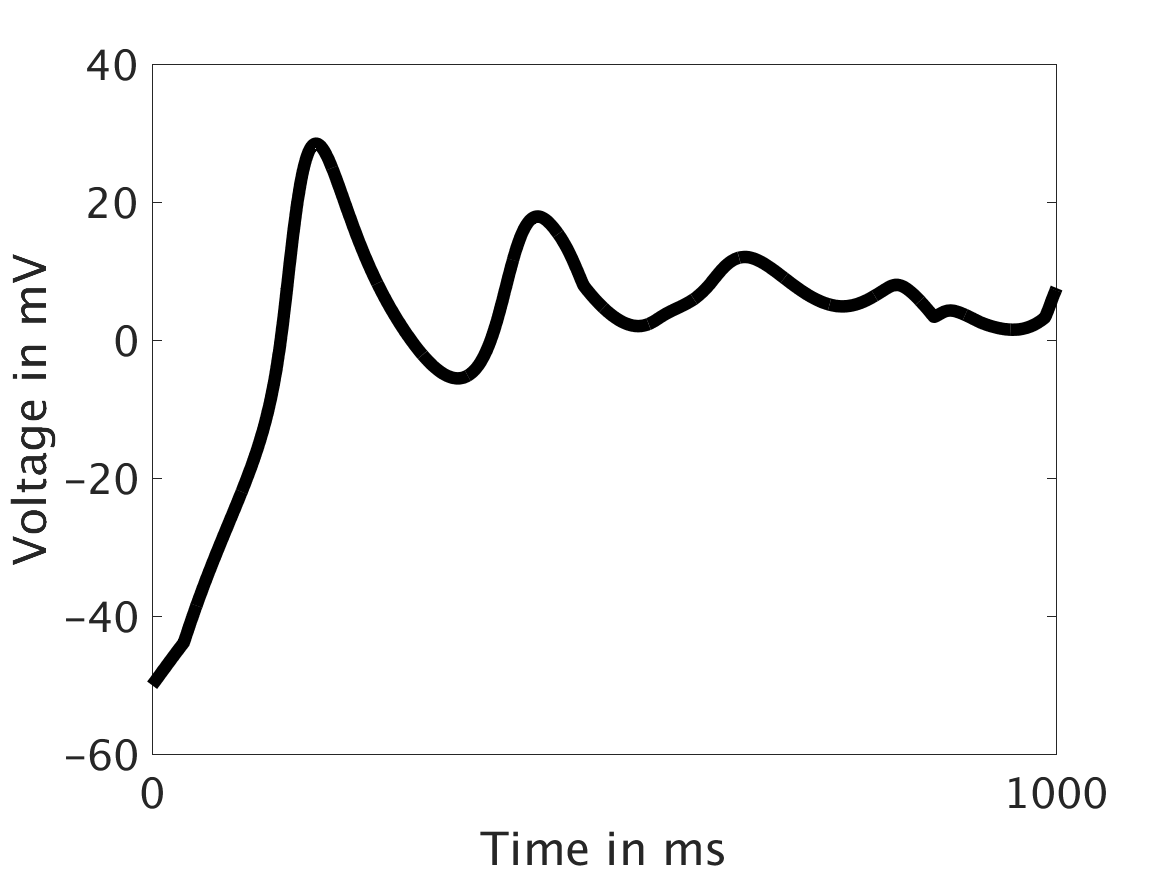
ω = 100:
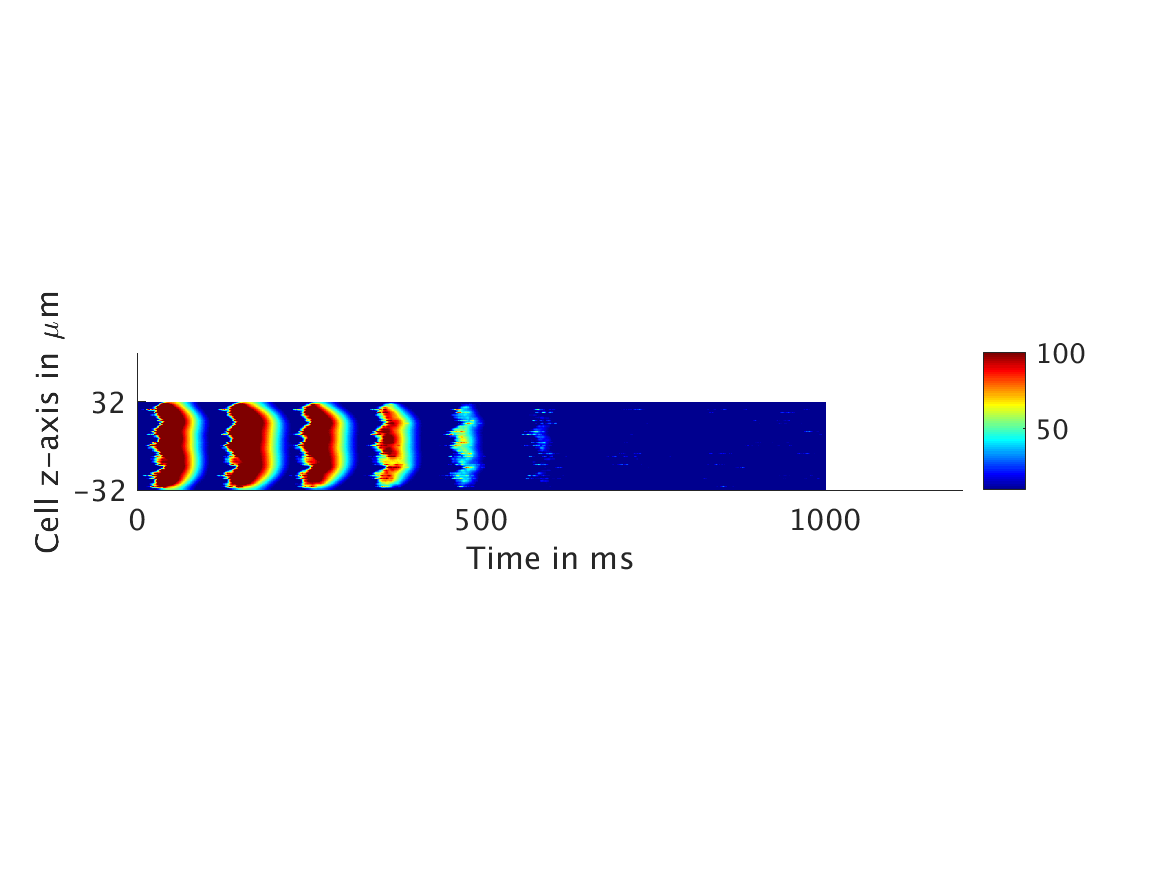
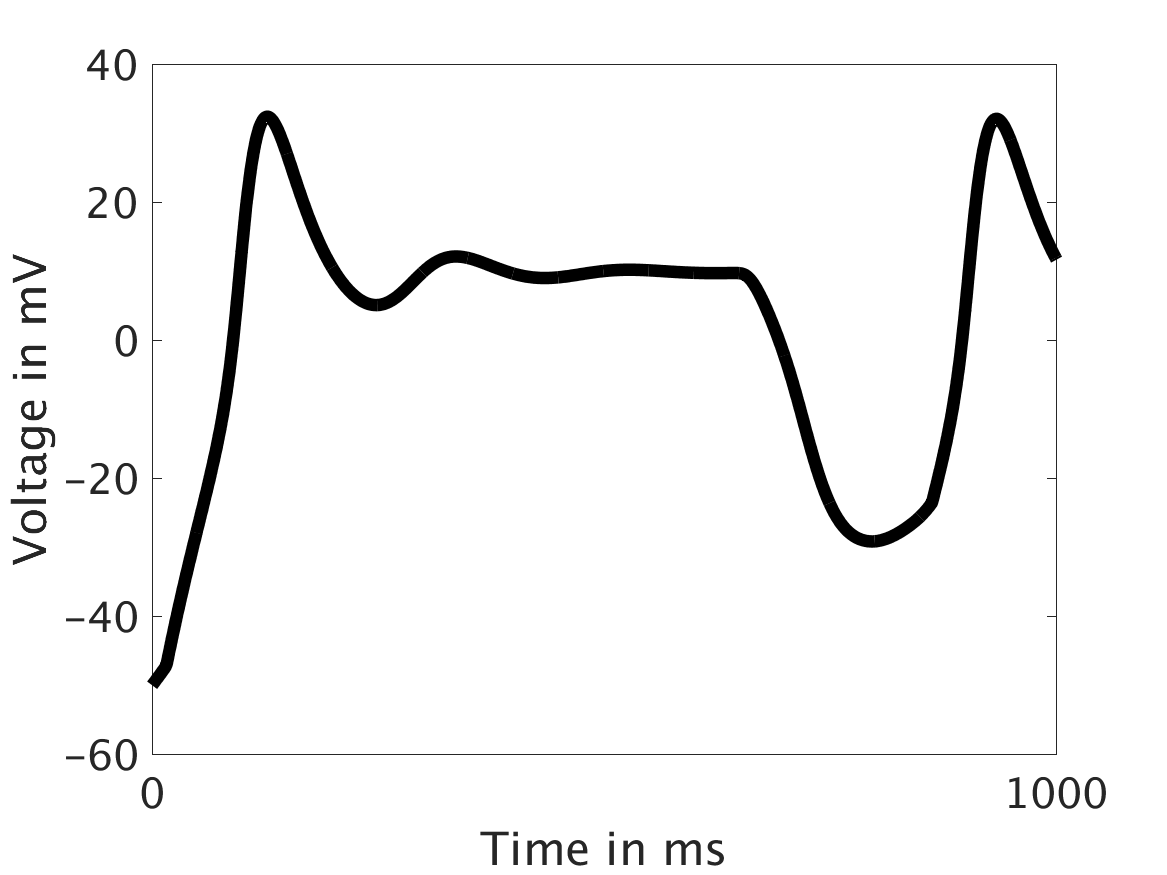
Conclusions and Future Directions
Our extended model completes the links between the electrical, calcium, and mechanical systems; simulations study coupling between the electrical and calcium systems. We introduce a calcium dependency into the voltage PDE, controlled by a scaling factor of feedback strength ω. This calcium dependency is the result of the calcium ion efflux from the cytosol out through the cell membrane, which generates a current with a significant effect on membrane potential, otherwise known as voltage. Our parameter study indicates that as ω increases, the effect on the voltage increases correspondingly, as is physiologically realistic: as the influence of the efflux current grows, the formerly steady periodicity of the voltage becomes more and more responsive to oscillations in the cytosol calcium concentration.
Further directions of this research would include implementation and parallelization of the third cytosol buffer species, the actin-myosin cross-bridges: while we have established the grounds for the behavior of this species mathematically, the relevant modifications have not yet been incorporated into our larger C code.
Links
Kallista Angeloff, Carlos Barajas, Alexander D. Middleton, Uchenna Osia, Jonathan S. Graf, Matthias K. Gobbert, and Zana Coulibaly. Coupling the Electrical and Calcium Dynamics in a Heart Cell. Technical Report HPCF-2016-15, UMBC High Performance Computing Facility, University of Maryland, Baltimore County, 2016. (HPCF machines used: maya.). Reprint in HPCF publications list
Poster presented at the Summer Undergraduate Research Fest (SURF)
Click here to view Team 1’s project
Click here to view Team 2’s project
Click here to view Team 3’s project
Click here to view Team 4’s project
Click here to view Team 6’s project
Click here to view Team 7’s project

